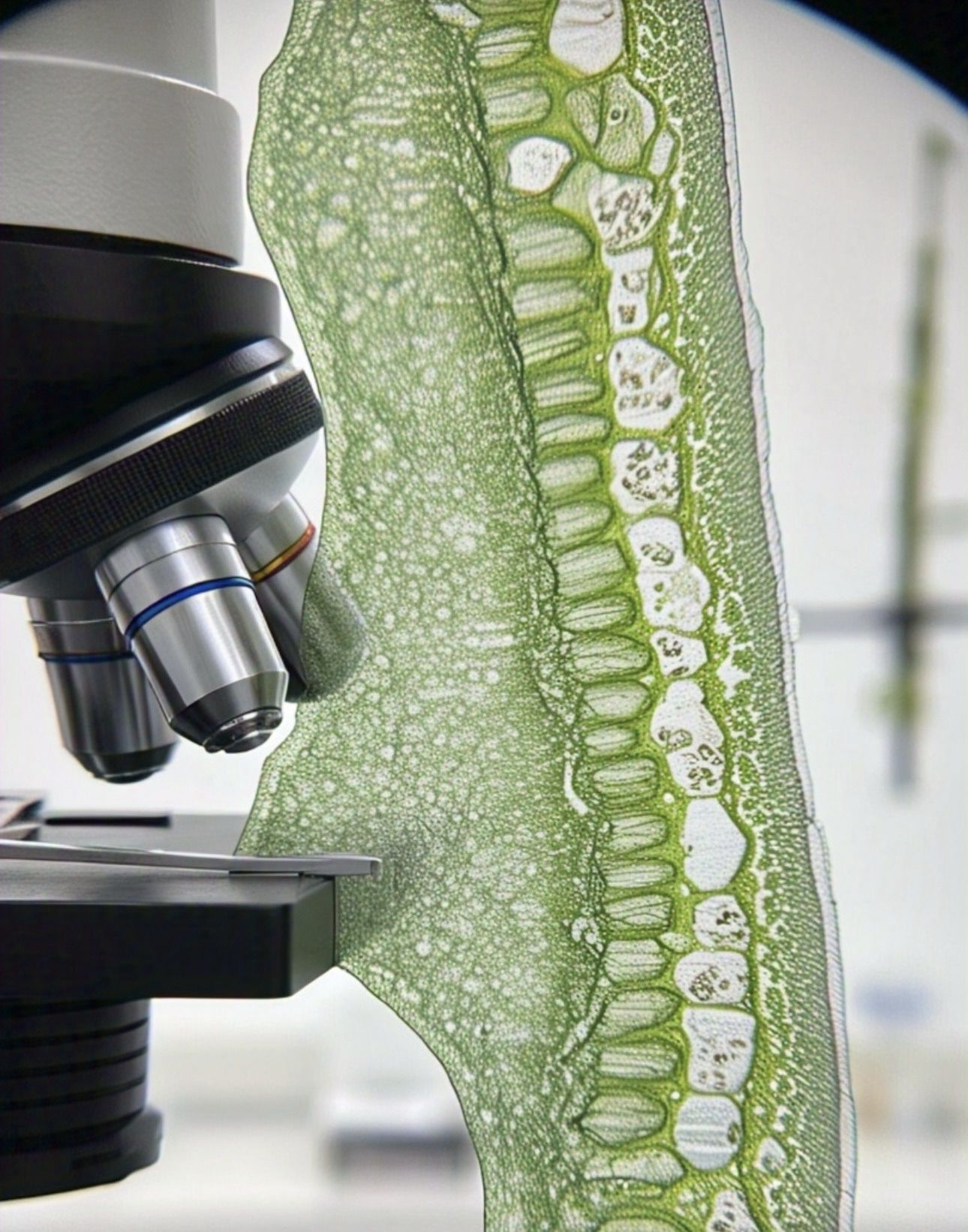
The plant world is composed of incredible intricacies — from the flow of nutrients through root hairs to the rapid movement of proteins across cellular membranes. Yet for centuries, scientists were limited in their ability to observe the minute details of these processes due to a fundamental physical limitation: the diffraction limit of light. Conventional optical microscopy could only resolve structures larger than 200 nanometers, leaving a vast realm of biological detail blurred or invisible.
Today, the emergence of super-resolution microscopy (SRM) is rewriting the rules of what we can observe in plant cells. With the ability to resolve structures as small as 20 nanometers — ten times finer than traditional methods — SRM is allowing researchers to explore the internal architecture of plant cells with unprecedented clarity. It is revolutionizing fields like plant developmental biology, cell signaling, cytoskeletal dynamics, and even molecular plant-pathogen interactions.
In this in-depth article, we explore how SRM works, its various types, key applications in plant biology, challenges, and the promising future it holds for plant research and precision agriculture.
The Diffraction Limit: A Barrier to Cellular Detail
In classical light microscopy, the resolution is restricted by the diffraction of light waves. According to the laws of optics, structures that are closer than approximately half the wavelength of the light used (about 200 nm for visible light) cannot be distinctly resolved. As a result, small structures such as protein complexes, organelle subdomains, and certain cytoskeletal elements appear fuzzy or merged in conventional microscopy.
This barrier has long constrained the visualization of dynamic and nanoscopic events within living plant cells — especially given the dense and complex nature of plant tissues. While electron microscopy provided high resolution, it required harsh fixation and could not be applied to live samples. A live, high-resolution alternative was desperately needed — and super-resolution microscopy delivered exactly that.
What Is Super-Resolution Microscopy?
Super-resolution microscopy refers to a family of advanced imaging techniques that surpass the diffraction limit, enabling visualization of biological structures with spatial resolutions down to 20–50 nanometers, and in some cases even finer. These techniques operate on principles such as spatial patterning of light, controlled activation of fluorophores, or the selective deactivation of fluorescence around a focal point. Crucially, SRM can be applied to living cells, enabling real-time observation of dynamic biological processes at the nanoscale.
In the context of plant biology, super-resolution microscopy allows scientists to examine subcellular organization, protein clustering, intracellular trafficking, and signaling pathways — areas that were once inaccessible with standard methods.
Major Super-Resolution Techniques in Plant Science
STED Microscopy (Stimulated Emission Depletion)
STED works by selectively depleting fluorescence around a central focal point using a doughnut-shaped laser beam. Only the center of the beam remains fluorescent, effectively reducing the point spread function and sharpening resolution.
In plants, STED microscopy has been particularly valuable in imaging actin filaments, microtubule arrays, and plasma membrane proteins. It has enabled high-speed imaging of cell plate formation during cell division, and visualization of protein nanoclusters involved in membrane trafficking. STED is compatible with live imaging, although it requires careful tuning to minimize phototoxicity in sensitive tissues.
SIM (Structured Illumination Microscopy)
SIM projects patterned light onto the sample and captures multiple images to generate interference patterns (Moiré patterns). These patterns are then computationally decoded to reconstruct a higher-resolution image — typically doubling the resolution of confocal microscopy.
SIM has proven ideal for imaging relatively thick or autofluorescent plant tissues, such as roots and leaves. Researchers have used SIM to explore chloroplast thylakoid structure, vesicle dynamics, and nuclear architecture. Since it uses lower light intensity compared to STED, SIM is suitable for long-term imaging of live tissues and organ development.
PALM and STORM (Localization Microscopy)
PALM (Photoactivated Localization Microscopy) and STORM (Stochastic Optical Reconstruction Microscopy) rely on the activation and precise localization of individual fluorescent molecules. By repeatedly imaging small subsets of fluorophores and computing their positions with nanometer precision, a super-resolved image is built over time.
In plant cells, PALM and STORM have enabled visualization of:
The organization of cellulose synthase complexes at the plasma membrane. The nanoscale distribution of hormone transporters such as the PIN family in the root apex. The structural remodeling of plasmodesmata in response to stress.
While the sample preparation for these techniques is more demanding, the payoff is ultra-high resolution — sometimes better than 20 nm — and detailed molecular mapping.
Breakthrough Discoveries Using Super-Resolution Microscopy
Super-resolution techniques have already transformed our understanding of several plant processes.
One major breakthrough has been the ability to visualize membrane nanodomains — small, specialized regions of the plasma membrane where signaling proteins cluster and interact. These structures, once only theorized, have now been clearly imaged in species such as Arabidopsis thaliana and Zea mays. They are now believed to play crucial roles in hormone perception, pathogen recognition, and membrane trafficking.
Another area that has greatly benefited from SRM is cytoskeletal dynamics. While actin filaments and microtubules are central to cell shape, polarity, and division, traditional methods failed to reveal the finer architecture and reorganization of these networks in real time. Super-resolution imaging has illuminated the subtle interactions between microtubules and organelles, as well as the mechanics of vesicle transport along cytoskeletal highways.
Furthermore, SRM has enabled the first detailed insights into the organization of cell walls at the nanoscale. By applying PALM/STORM, researchers can now observe how cellulose microfibrils are arranged, how they respond to stress or hormone treatment, and how cell wall enzymes are spatially distributed during tissue remodeling.
Challenges in Adapting SRM to Plant Systems
Despite its remarkable capabilities, super-resolution microscopy comes with technical and biological challenges, particularly in plant tissues.
Plant cells often produce strong autofluorescence due to pigments like chlorophyll and lignin, which can interfere with fluorophore detection. Moreover, plant tissues tend to be thicker and more optically dense than animal cells, making imaging deeper layers a challenge. The presence of the cell wall, although biologically crucial, adds another barrier to imaging.
Another significant limitation is labeling efficiency. In plant systems, stable transformation and reporter gene expression are not always straightforward. While Arabidopsis is a model system well-suited for genetic labeling, many crop species are less amenable to transformation and thus harder to study using SRM.
To overcome these challenges, researchers are adopting strategies like:
Tissue clearing methods to reduce opacity. Adaptive optics to correct light distortion. Development of plant-specific fluorescent proteins with reduced photobleaching. Optimized sample mounting and imaging chambers for live-cell compatibility.
Future Directions: Where is Super-Resolution Microscopy Headed in Plant Research?
The future of SRM in plant biology is incredibly promising. Already, we are seeing its integration with other powerful technologies such as machine learning, computational modeling, and biosensor development.
In particular, deep learning algorithms are being trained to enhance image reconstruction, automate feature recognition, and even predict the future behavior of cells based on live imaging data.
Expansion microscopy is another exciting development, where plant tissues are physically expanded using polymers, allowing conventional microscopes to achieve SRM-level resolution without the need for high-powered lasers.
Integration with correlative microscopy, where SRM is combined with techniques like electron microscopy (CLEM), will further bridge the gap between molecular localization and ultrastructural context.
Additionally, SRM is beginning to play a role in plant pathology and stress biology. Scientists are exploring how pathogen effector proteins reorganize plant cytoskeletal structures or membrane domains — insights that could lead to more targeted disease resistance strategies.
Conclusion
Super-resolution microscopy has opened a new era in plant biology — one where the inner world of plant cells is no longer a mystery veiled in blur. By breaking the limits imposed by light diffraction, this technology is allowing researchers to visualize the nanoscopic details of life: the organization of proteins, the dynamics of the cytoskeleton, the architecture of membranes, and the pathways of intracellular signaling.
As tools continue to evolve and accessibility improves, SRM will become increasingly central to answering the most pressing questions in plant science — from how roots navigate soil microenvironments, to how leaves adapt to climate change, and how crops respond to pathogens at the cellular level.
References
Komis, G., Šamajová, O., Ovečka, M., & Šamaj, J. (2015). Super-resolution microscopy in plant cell imaging. Trends in Plant Science, 20(12), 834–843. https://doi.org/10.1016/j.tplants.2015.08.004
Mota, A., Vavrdová, T., Ovečka, M., Šamajová, O., & Šamaj, J. (2021). Advanced microscopy approaches to study plant cell walls. Frontiers in Plant Science, 12, 651133. https://doi.org/10.3389/fpls.2021.651133
Schubert, V. (2017). Super-resolution microscopy–Applications in plant cell research. Frontiers in Plant Science, 8, 531. https://doi.org/10.3389/fpls.2017.00531
Ries, J., Kaplan, C., Platonova, E., Eghlidi, H., & Ewers, H. (2012). A simple, versatile method for GFP-based super-resolution microscopy via nanobodies. Nature Methods, 9, 582–584. https://doi.org/10.1038/nmeth.1991
Ursache, R., Andersen, T. G., Marhavý, P., & Geldner, N. (2018). A protocol for combining fluorescent proteins with histological stains for diverse cell wall components. The Plant Journal, 93(2), 399–412. https://doi.org/10.1111/tpj.13784
Ovečka, M., Vaškebová, L., Komis, G., Luptovčiak, I., Smertenko, A., & Šamaj, J. (2015). Multiscale imaging of plant development by light-sheet fluorescence microscopy. Nature Plants, 1(6), 15110. https://doi.org/10.1038/nplants.2015.110