Plants may appear simple from the outside as a collection of leaves, stems, flowers and roots but inside every plant lies a world of extraordinary complexity. Each organ contains thousands of cell types, and every single cell has a unique job, identity, and history. For decades, plant scientists could only study this complexity by measuring gene expression in bulk tissues, blending millions of cells together and averaging their signals. It was like listening to a choir without ever hearing the individual voices.
But the rise of single-cell transcriptomics has changed everything. This cutting-edge technology lets researchers zoom into individual cells and measure which genes are active in each one. Instead of treating a leaf or root as a uniform tissue, scientists can now uncover the hidden diversity of cell types, how they communicate, and how they respond differently to stress.
Single-cell transcriptomics is rewriting plant physiology at its core. It has revealed unknown cell populations, uncovered stress-responsive hotspots, transformed our understanding of development, and opened new possibilities for crop improvement. In this article, we explore what single-cell transcriptomics is, why it matters for plant physiology, and how it is shaping the future of climate-resilient agriculture.
What Is Single-Cell Transcriptomics and Why Does It Matter?
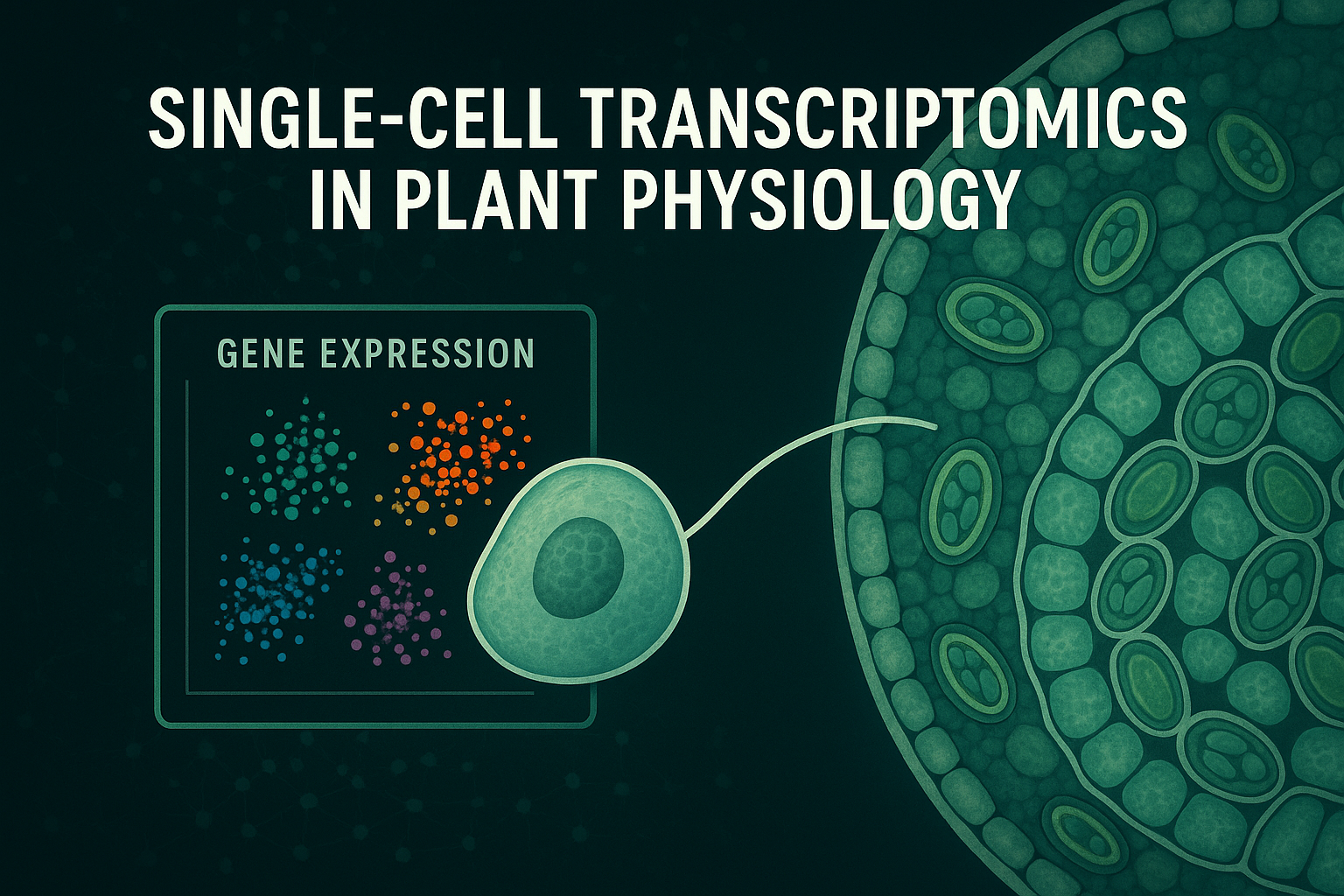
Traditional gene expression studies measure mRNA from thousands or millions of cells simultaneously. While powerful, this approach hides differences between cells. If only a small group of cells expresses a stress-responsive gene, bulk sequencing may fail to detect it. If two cell types respond very differently to drought or salinity, their opposing signals may cancel out in an average.
Single-cell transcriptomics solves this problem by isolating individual cells and sequencing their mRNA one cell at a time. The result is a “gene expression fingerprint” for each cell, providing unprecedented detail about cellular identity and function.
This matters tremendously for plant physiology because plants are highly heterogeneous organisms. A single root contains epidermal cells, cortex cells, endodermis, pericycle, vascular initials, xylem, phloem and dozens of transitional cell types. Each responds differently to nutrients, microbes, drought, or mechanical stress. Single-cell transcriptomics allows us to see this complexity clearly for the first time.
Understanding Tissues Cell by Cell
One of the most exciting contributions of single-cell transcriptomics has been the discovery of new or rare plant cell types. Earlier methods could not detect cell populations that made up less than 1% of a tissue. But with single-cell approaches, even tiny cell groups become visible.
In roots, single-cell studies have identified previously unknown subtypes of cortex cells that activate nutrient transporters under stress. In leaves, researchers have detected specialized guard cell precursors, mesophyll subpopulations with distinct photosynthetic profiles, and vascular cells that control long-distance signaling. These discoveries change how we interpret plant physiology at the tissue level because each newly identified cell type may have unique metabolic or regulatory roles.
Another major insight is that plant cells are not fixed in identity. Single-cell transcriptomics has shown that many cells exist in transitional states, especially during growth, regeneration, and stress. These transitional states are invisible in bulk studies, yet they are critical for understanding how plants reshape tissues after injury, respond to fluctuating environments, or allocate resources during development.
This cellular-resolution map of plant tissues allows physiologists to build more accurate models of water flow, photosynthesis, nutrient uptake, and hormonal communication.
How Single Cells React to the Environment
Plants face many stresses like heat, drought, salinity, pathogens and nutrient limitations. Traditional studies often described stress responses as tissue-wide events, but single-cell transcriptomics shows that stress sensing and reaction begin in specific cell groups long before the entire tissue responds.
Under drought, for example, the earliest transcriptomic changes often occur in guard cells and xylem-adjacent tissues. These cells activate ABA signaling, osmotic adjustment genes and protective proteins earlier than neighboring cells. Similarly, during salt stress, endodermal and pericycle cells show unique ion transport signatures that are diluted in bulk analysis.
Single-cell studies have also revealed that some cells act as “stress coordinators.” These rare cells may detect environmental changes and send signals to surrounding tissues via hormones or mobile peptides. Identifying these coordinator cells offers new opportunities to engineer stronger stress tolerance by targeting very specific cell populations.
Pathogen responses are even more striking. When a plant detects a pathogen, only a small cluster of cells initiates immune signaling, while others remain stable to maintain tissue function. Without single-cell data, this fine-scale pattern would remain hidden, leaving us with an oversimplified picture of immunity.
By uncovering the cell-specific architecture of stress responses, single-cell transcriptomics is guiding the development of crops that react faster and more efficiently to the environment.
Building Climate-Resilient Crops
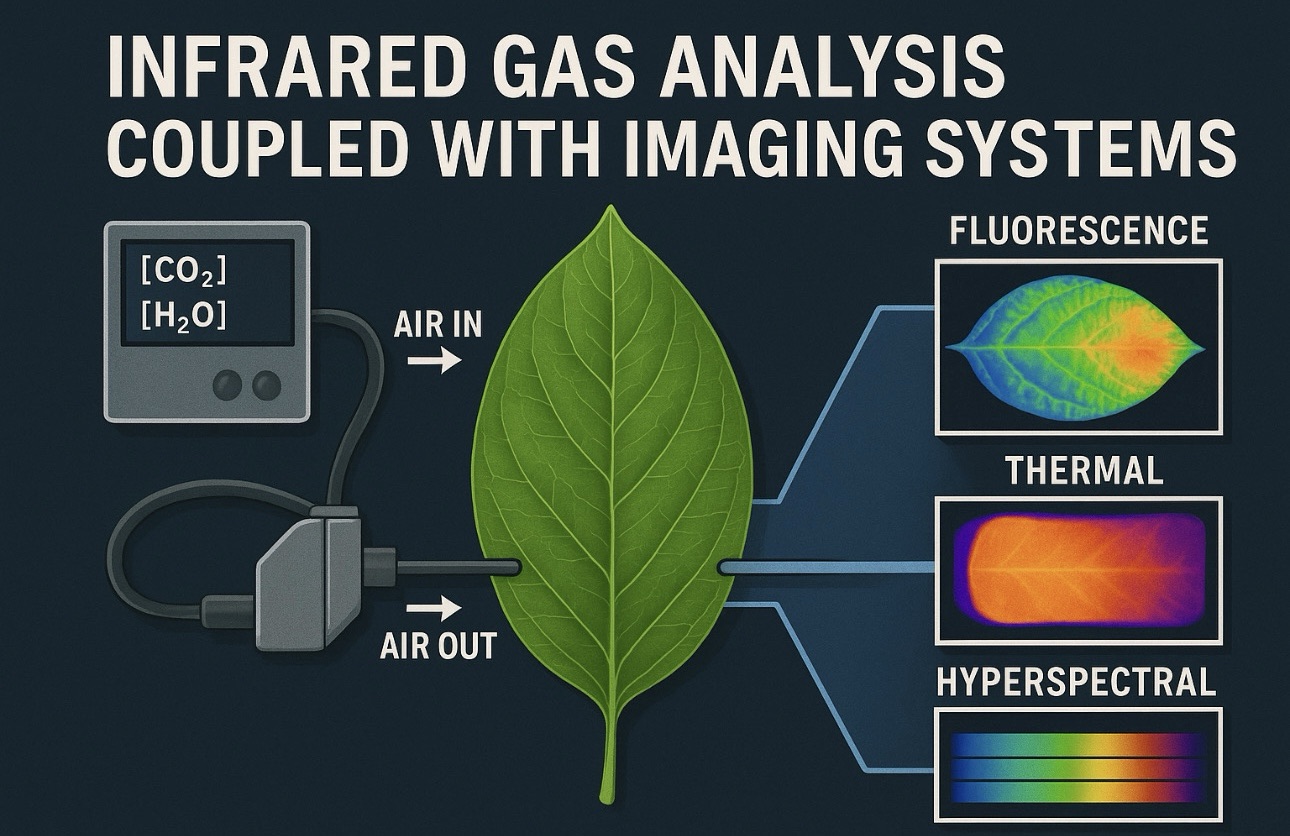
The potential of single-cell transcriptomics for crop improvement is immense. One major application is identifying cell types that limit photosynthesis or stress tolerance. For example, leaf mesophyll cells differ in their CO₂ assimilation capacity, and single-cell data helps pinpoint which subpopulations contribute most to performance under heat or drought.
Root architecture is another major target. Single-cell approaches reveal how root tips generate new xylem vessels during drought, how endodermal cells build stronger barriers, and how specific cortical cells support beneficial microbes. These insights can guide breeding programs to enhance water use efficiency, nutrient uptake, and resilience.
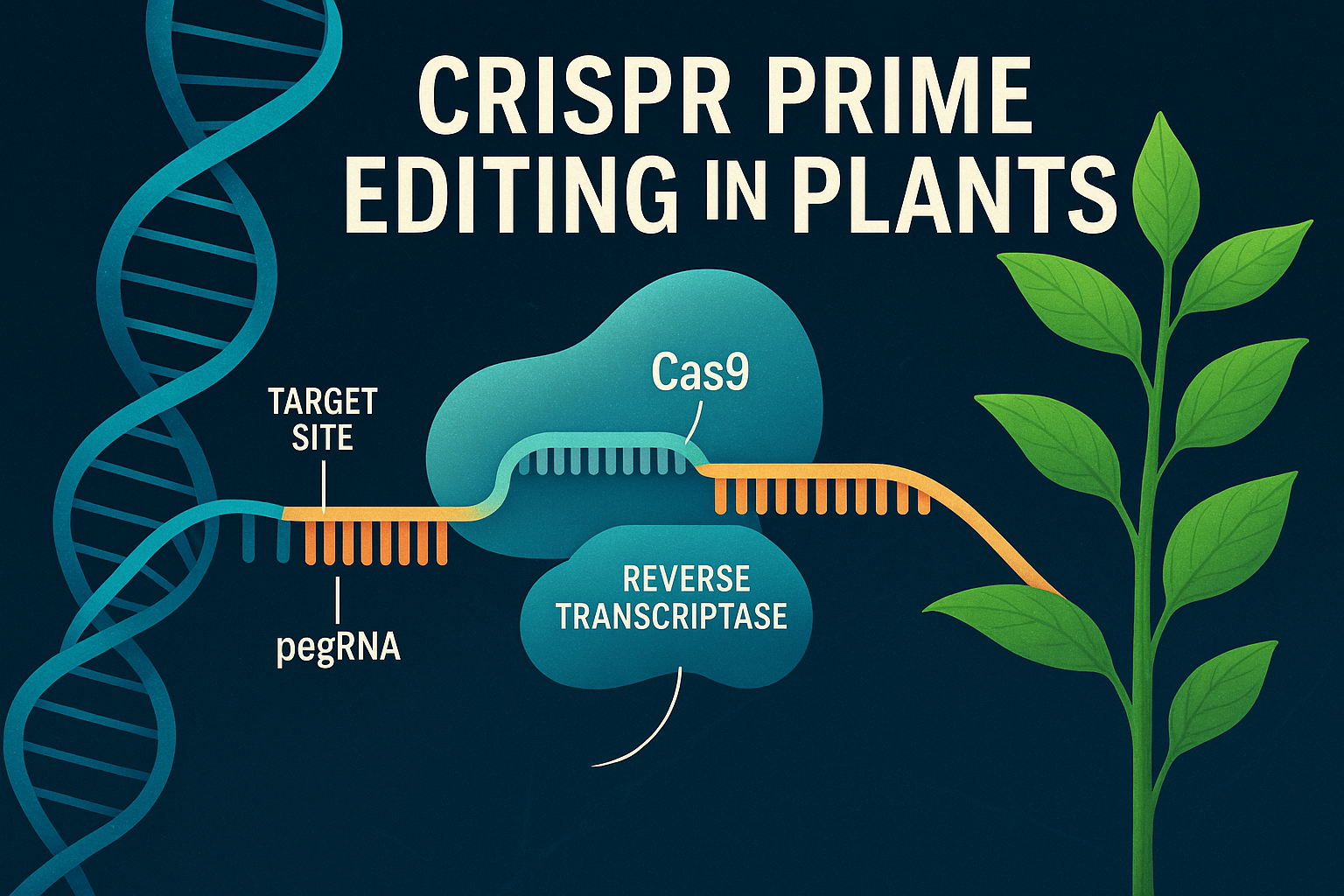
Single-cell transcriptomics is also accelerating gene discovery. Genes that are highly expressed only in rare or transient cell types would otherwise be missed. Identifying these “hidden regulators” allows scientists to manipulate key developmental or stress pathways with precision.
Future technologies will take this further. Single-nucleus sequencing, spatial transcriptomics, and cell-specific epigenomics are emerging tools that combine gene expression with spatial and regulatory information. Instead of separating cells, spatial methods preserve tissue structure, showing exactly where each cell type sits and how signals move across tissues.
Together, these tools will create a multi-layered map of plant physiology, from gene expression to cell function to whole-plant behavior. Climate-resilient crops will be designed not only by modifying whole tissues but by targeting individual cells that drive adaptation.
Conclusion
Single-cell transcriptomics has given plant science a new lens, one powerful enough to reveal the hidden diversity, dynamics and decision-making inside plant tissues. Instead of viewing leaves and roots as uniform units, we can now see them as intricate mosaics of highly specialized cells, each responding to the environment in its own way.
This new perspective is already reshaping plant physiology. It helps explain how plants grow, how they sense stress, and how they coordinate complex responses across tissues. As technology advances, single-cell tools will become even more essential for building the next generation of crops that are more productive, more resilient, and better adapted to the challenges of a changing climate.
Plants have always been complex, but now, for the first time, we can truly see that complexity