Plants are made up of an extraordinary diversity of cells, each performing highly specialized functions. A guard cell regulates gas exchange, a mesophyll cell drives photosynthesis, a root hair absorbs nutrients, and a vascular cell transports water or sugars. Yet, for a long time, plant physiology treated tissues as uniform units, averaging signals across thousands or millions of cells. While this approach provided valuable insights, it masked the true cellular diversity that underpins plant function. The emergence of single-cell proteomics is transforming this perspective. Instead of measuring proteins at the tissue or organ level, scientists can now examine the protein composition of individual plant cells. This shift is profound because proteins—not genes carry out most physiological processes. By studying proteins at single-cell resolution, researchers are uncovering how different cells respond uniquely to stress, development, and environmental signals. Single-cell proteomics opens a new window into plant physiology, revealing hidden heterogeneity, dynamic regulation, and cell-specific stress strategies that were previously invisible. As climate challenges intensify, understanding how individual cells cope with stress may hold the key to designing resilient crops.
Why Proteins Matter at the Single-Cell Level
Genes provide the blueprint, but proteins are the active machinery of life. Enzymes catalyze reactions, transporters move ions and metabolites, receptors perceive signals, and structural proteins shape cells. Importantly, protein abundance, modification, and activity do not always correlate directly with gene expression. Two cells with similar transcript profiles may behave very differently because of differences in protein stability, localization, or post-translational modifications. In plant tissues, cellular heterogeneity is especially pronounced. Neighboring cells often experience different microenvironments, light intensities, water availability, and mechanical forces. Under stress, some cells activate protective pathways rapidly, while others respond more slowly or not at all. Bulk proteomic approaches average these differences, obscuring critical information. Single-cell proteomics allows researchers to capture this diversity. It reveals which proteins are present in each cell, how their abundance changes over time, and how cells reprogram their proteome under stress. This approach is particularly valuable in plant physiology, where responses are often spatially organized for example, stress perception in roots versus signal execution in shoots, or differential responses across leaf layers. By focusing on proteins rather than transcripts, single-cell proteomics provides a more direct view of physiological function, bridging the gap between molecular regulation and whole-plant behavior.
Technological Advances Driving Single-Cell Proteomics in Plants
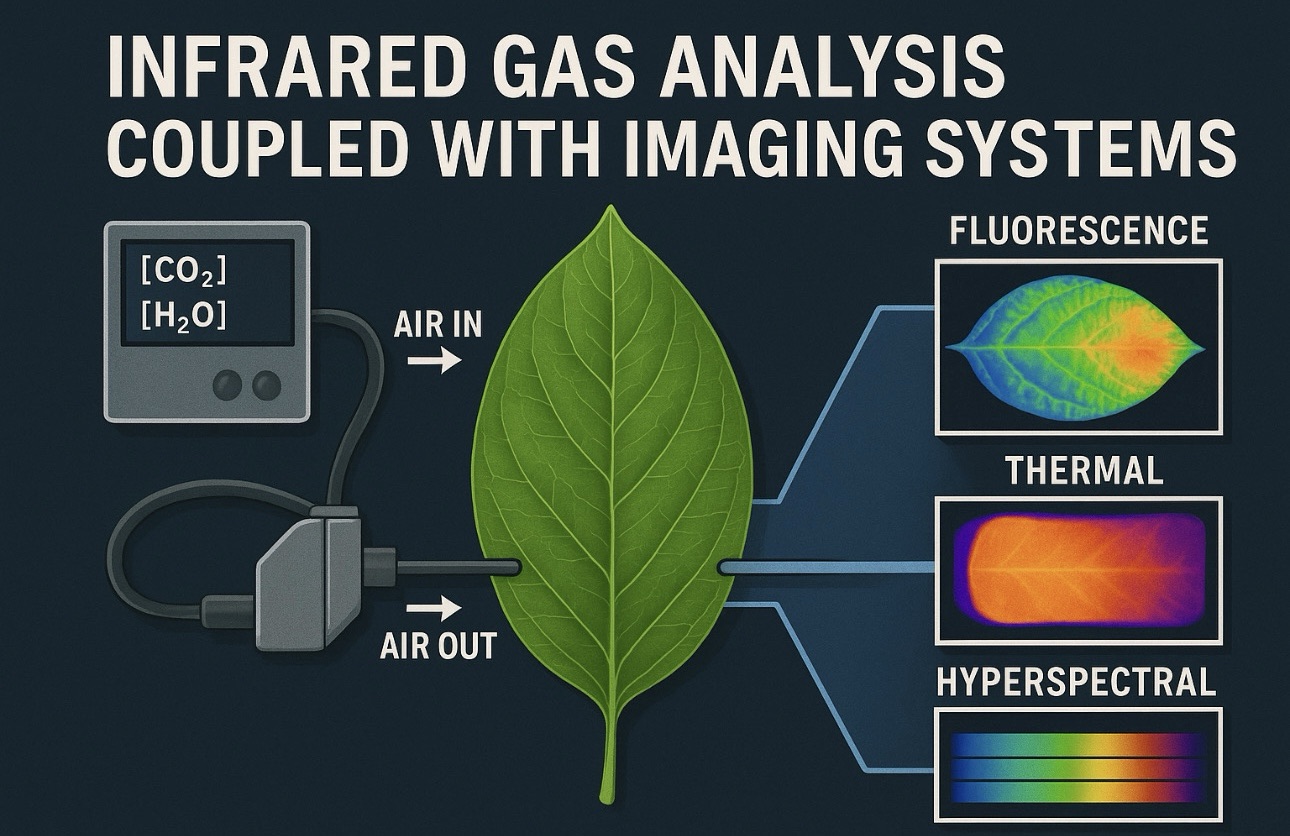
Applying single-cell proteomics to plants presents unique challenges. Plant cells are encased in rigid cell walls, contain large vacuoles, and exhibit strong metabolic compartmentalization. Isolating intact single cells or protoplasts without altering protein composition requires careful optimization. Recent technological breakthroughs have made this possible. Advanced cell isolation methods, including gentle protoplasting, microdissection, and microfluidic platforms, now allow individual plant cells to be captured with minimal perturbation. Ultra-sensitive mass spectrometry has dramatically improved protein detection limits, making it feasible to analyze the tiny protein quantities present in a single cell. Innovations such as nano-scale liquid chromatography, optimized sample preparation workflows, and multiplexing strategies enable hundreds or even thousands of proteins to be quantified from individual cells. In parallel, improvements in bioinformatics allow complex datasets to be interpreted, clustering cells based on proteomic profiles and identifying cell-specific pathways. Single-cell proteomics is often combined with other approaches, such as single-cell transcriptomics or imaging, to provide a multi-layered view of cellular regulation. This integrated strategy is particularly powerful for plant physiology, where cellular identity, metabolic state, and environmental context are tightly intertwined.
Stress Responses at Cellular Resolution
One of the most exciting applications of single-cell proteomics is the study of plant stress responses. Environmental stresses rarely affect all cells equally. In a drought-stressed root, some cells experience severe dehydration while others remain relatively protected. In a heat-stressed leaf, cells near the surface may overheat while deeper layers remain cooler. Single-cell proteomics reveals how different cells deploy distinct stress strategies. Some cells increase antioxidant enzymes to manage oxidative stress. Others adjust ion transporters to maintain osmotic balance. Certain cells upregulate chaperones and proteases to protect protein structure, while others shift metabolism to conserve energy. This cellular-level insight helps explain why plants can survive stress even when some cells are damaged. It also identifies key cell types that act as stress sensors or signal hubs. For example, specific root cells may initiate drought signaling, while vascular cells transmit the signal throughout the plant. Single-cell proteomics has also shed light on immune responses. During pathogen attack, only a subset of cells may activate defense pathways initially. By capturing proteomic changes in these cells, researchers can understand how immune signals spread and how localized responses become systemic.
Implications for Development, Breeding, and Climate Resilience
Beyond stress physiology, single-cell proteomics is reshaping our understanding of plant development. Developmental transitions such as root branching, leaf differentiation, and flowering are driven by precise changes in protein networks within specific cells. Single-cell proteomics allows these transitions to be mapped with unprecedented resolution. For crop breeding, this technology offers powerful opportunities. Stress tolerance often depends on the behavior of particular cell types rather than the whole plant. By identifying proteomic signatures associated with resilient cells, breeders can target traits more precisely. This could lead to crops that respond faster to stress, maintain function under extreme conditions, or recover more effectively after damage.
Single-cell proteomics also supports the development of cell-type–specific engineering strategies. Instead of modifying genes globally, future approaches may target specific cells, such as guard cells or root cortex cells, optimizing their proteome for stress resilience without compromising overall growth. As analytical tools continue to improve, single-cell proteomics is likely to become a cornerstone of systems-level plant physiology, integrating molecular, cellular, and whole-plant perspectives.
Conclusion
Single-cell proteomics represents a paradigm shift in plant science. By moving beyond tissue averages and focusing on individual cells, it reveals the true complexity of plant physiology. Proteins emerge as dynamic regulators that differ from cell to cell, shaping how plants grow, adapt, and survive. In a future defined by climate uncertainty, understanding plant responses at the finest possible resolution will be essential. Single-cell proteomics offers exactly that a way to see how each cell contributes to resilience, one protein at a time. As this technology matures, it promises to deepen our understanding of plant life and guide the development of crops that are smarter, stronger, and better adapted to a changing world.